Tiberiu Teșileanu a physicist working in neuroscience
Spring simulation
 |
masses have different sizes |
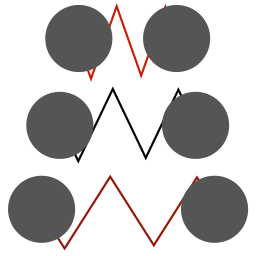 |
spring stretch indicated by redness |
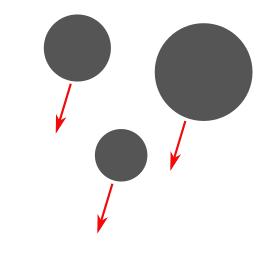 |
uniform acceleration (change using arrow in the corner) |
 |
dots move at constant velocity, except for reflections at walls |
 |
masses slowed down by air resistance |
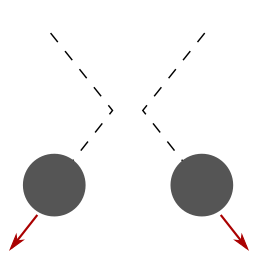 |
collisions between masses are perfectly elastic |
 |
collisions with walls lose energy |
I’m a Research Software Engineer in the CTRL team at Meta.
Use the navigation menu at the top/left to learn more, or contact me using one of the links below.